Method Developed by Children’s Hospital Los Angeles Researchers Automates Brain Cell Mapping
Neuroscience graduate students at Children’s Hospital Los Angeles have developed an automated method that could save time and work for laboratories around the country by streamlining the process of identifying and mapping brain cells. Scientists want to understand how brain cells develop over time because the way these cells, called neurons, develop influences how they function, or how they malfunction in neurodevelopmental disorders.
To understand how children’s brains develop, and how this development goes awry in disorders such as autism, neuroscience researchers catalog various brain cells by their function and where they are located in tissues. Mapping these cells can help scientists understand how different types of neurons interact in networks to direct brain functions. This basic research could eventually be used to guide therapies for various neurodevelopmental disorders.
“At least when it comes to the brain, there is a spatial organization,” says Ramin Ali Marandi Ghoddousi, a PhD candidate in neuroscience who researches early brain and motor development in the Levitt Laboratory. “Certain cell types reside in one part of the tissue and other cell types in other parts. Sometimes they are interspersed with each other. But where they are found in the tissue implicates the different neuron functions.”
“There are tens of thousands of different cell types,” says Pat Levitt, PhD, Senior Vice President, Chief Scientific Officer and Director of The Saban Research Institute at Children’s Hospital Los Angeles. “Which kind of neurons in the brain are expressing certain genes? In the brain, anatomy is everything.”
Aside from location, scientists also categorize cells by the genes they express -- in particular, their messenger RNA (mRNA), which carries genetic information from the DNA in a cell’s nucleus to its cytoplasm where the instructions are read and translated into the proteins that direct the cell’s development and function.
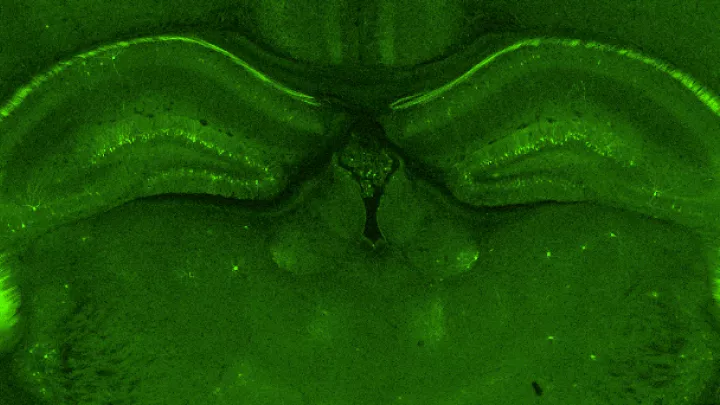
Researchers use a standard method to identify cells, called multiplex fluorescent in-situ hybridization, or mFISH. This technique stains the mRNA within cells with a fluorescent probe that induces the genetic material in the cells to light up under a microscope--rather like a scattering of holiday lights. The illuminated mRNA can help identify the type of cell. Some commercial multiplexing methods can characterize up to 40 genes at a time in a tissue section.
To distinguish whether a bit of mRNA belongs to one cell versus another in a tissue section, researchers perform a second staining procedure that marks the nucleus of each cell with a separate fluorescent signal, so any mRNA molecules that overlap with a particular cell nucleus will get assigned to that cell.
However, using a standard nuclear stain makes it more difficult to see where one cell ends and the other begins. “If you are staining that neuron, you actually can’t see the boundaries,” says Dr. Levitt, who also holds the Simms/Mann Chair in Developmental Neurogenetics. “It is like you can identify the house, but you can’t identify the street.” It is necessary to define the cell boundaries to assign the mRNA expression to the appropriate cell. “You go in and circle it manually,” says Dr. Levitt, “Imagine doing that for thousands of cells.”
The origin of SCAMPR
Back in the early days of the COVID-19 pandemic, when much of the world including scientific laboratories, began to shut down, Dr. Levitt discussed with his students some potential research projects that they could perform remotely. “Ramin spoke up and said he had been generating this data on brain structure,” recalls Dr. Levitt.
After two years of zoom calls and teaching themselves to code in R, a statistical programming language, Ali Marandi Ghoddousi and his fellow students developed a technique that could streamline the painstaking work of identifying, mapping, and analyzing cells. Dubbed SCAMPR— for single-cell automated multiplex pipeline for RNA quantification and spatial mapping— this method automates the identification of different cell types by analyzing their levels of gene expression and where they are located in a tissue section.
“SCAMPR sprang out of necessity, essentially,” says Ali Marandi Ghoddousi. “We used these multiplex fluorescent in-situ techniques in our lab for our experiments and realized that there wasn’t really a good method to analyze the data that we were getting back.” The method that the students created could speed up research by taking out the manual labor involved. “With SCAMPR, it is more than 90% automated,” notes Dr. Levitt.
Measuring mRNA
SCAMPR can also detect multiple genes that might appear in a cell together. “We will be able to measure gene expression to determine what kind of cell it is, and also where the gene expression occurs during our experiments,” says Ali Marandi Ghoddousi.
A few automated programs can do this as well, but most look for the mRNA expressed in the cell nucleus. However, the bulk of a cell’s mRNA is found in the cell cytoplasm—the watery internal part of the cell. “With SCAMPR we can identify the mRNA inside the nucleus as well as inside the rest of the cell,” says Ali Marandi Ghoddousi. “Before the invention of SCAMPR, people had attempted to do this by either guessing the boundaries of the cell or using the nucleus of the cell as a starting point and expanding out to estimate the boundary of the entire cell.”
SCAMPR was developed by Ali Marandi Ghoddousi, with Valerie M. Magalong, then a research assistant, with the participation of Anna Kamitakahara, PhD, Assistant Professor of Research at the Keck School of Medicine of USC. According to Ali Marandi Ghoddousi, SCAMPR can be useful for the average lab that wants to experiment with multiplex or high-dimensional in-situ hybridization but doesn’t have access to huge computational power. Some labs around the country have already contacted him expressing interest in the method. A paper describing SCAMPR was published in Cell Reports Methods.