Genetics and Environment Impact Familial Depression
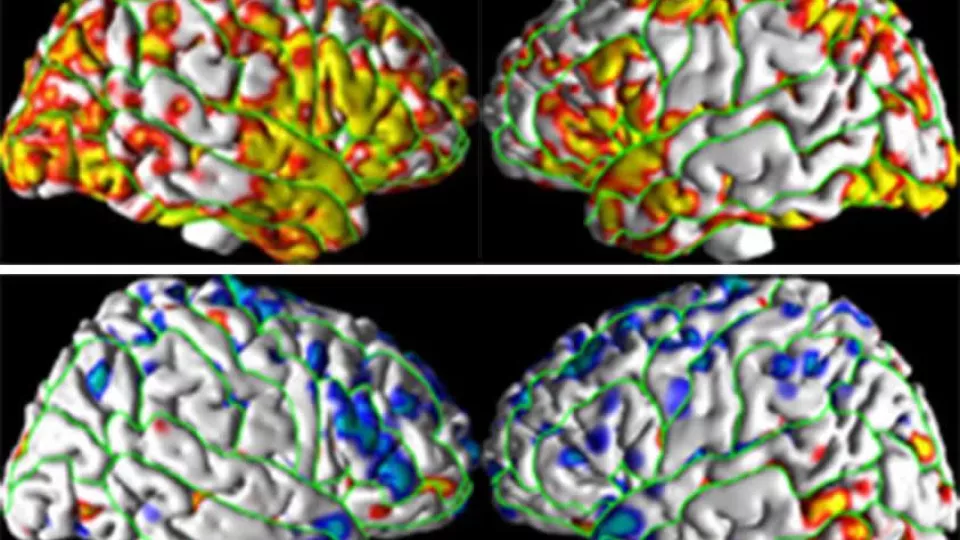
A genetic variation, or the “s” allele, was associated with thicker cortex in the high risk (top row) but with thinner cortex in the low risk (bottom row) individuals. Image courtesy of Ravi Bansal, PhD, Children’s Hospital Los Angeles.
Building on a 30-year, three-generation study of depressed individuals, their children and offspring, a study published in the journal Psychiatry Research: Neuroimagingprovides a better understanding of the familial risk for depression and the role neuroplasticity might have in increasing the risk of developing depression.
The research by scientists at Children’s Hospital Los Angeles and Columbia University shows a link between a particular allele for the serotonin found at a higher frequency in those at risk of depression because of family history, and those who go on to develop major depressive disorder.
Prior work by Bradley S. Peterson, MD, director of the Institute for the Developing Mind (IDM) at Children’s Hospital Los Angeles, and colleagues showed that individuals at familial risk for depression – whether or not they have exhibited signs of depression in the past – have a pattern of thinning in certain cortical regions of the brain. Additional brain loss occurs in other areas of the brain in those who do manifest depressive symptoms.
Several genes have been associated with depression, especially those genes that alter serotonin signaling. Serotonin is a chemical messenger in the brain that acts as a mood stabilizer. A genetic variation, or allele, found in the promoter region of the serotonin transporter was found to affect this brain loss.
The new study found that among those at low risk for depression based on family history, the “S” (short) allele is associated with a thinner cortex than those with the “L” (long) allele. However, in the individuals at high risk for depression, the correlation is actually reversed and more cortical thinning is found in those with the “L” allele.
In addition, those with the “S” allele were at higher risk of developing depression than those with the “L” allele when exposed to the same type and amount of stressors.
“These findings suggest that brain plasticity also plays a role in depression,” said first author Ravi Bansal, PhD, researcher at the IDM at Children’s Hospital Los Angeles and professor of pediatrics at Keck School of Medicine of the University of Southern California. “It could be that those with the “L” allele or genetic variation can better adapt to their environment. Those with the “S” allele appear to have lower brain plasticity and perhaps have fewer compensatory factors in the brain that allow them to adapt to adverse conditions.”
The study looked at brain MR images of 120 individuals, those at both high and low familial risk of depression, along with genotype data. Further scientific studies looking at cortical thickness, combined with genotyping, could also have the potential to predict who will develop depression and how individuals respond to anti-depressant medications, Bansal said.
This work was partially supported by the National Institute of Mental Health (grant #MH36197) and the Silvio O. Conte Center (grant #1P50MH090966).